About HitAnno
HitAnno is a scalable model built on a hierarchical transformer architecture for accurate cell type annotation on large scATAC-seq datasets. HitAnno constructs “cell sentences” by leveraging accessibility profiles on cell-type-specific peaks, capturing the epigenomic cell heterogeneity. The model adopts a two-level attention mechanism to capture both peak-level and peak-set-level dependencies, enabling hierarchical feature integration for reliable annotation performance.
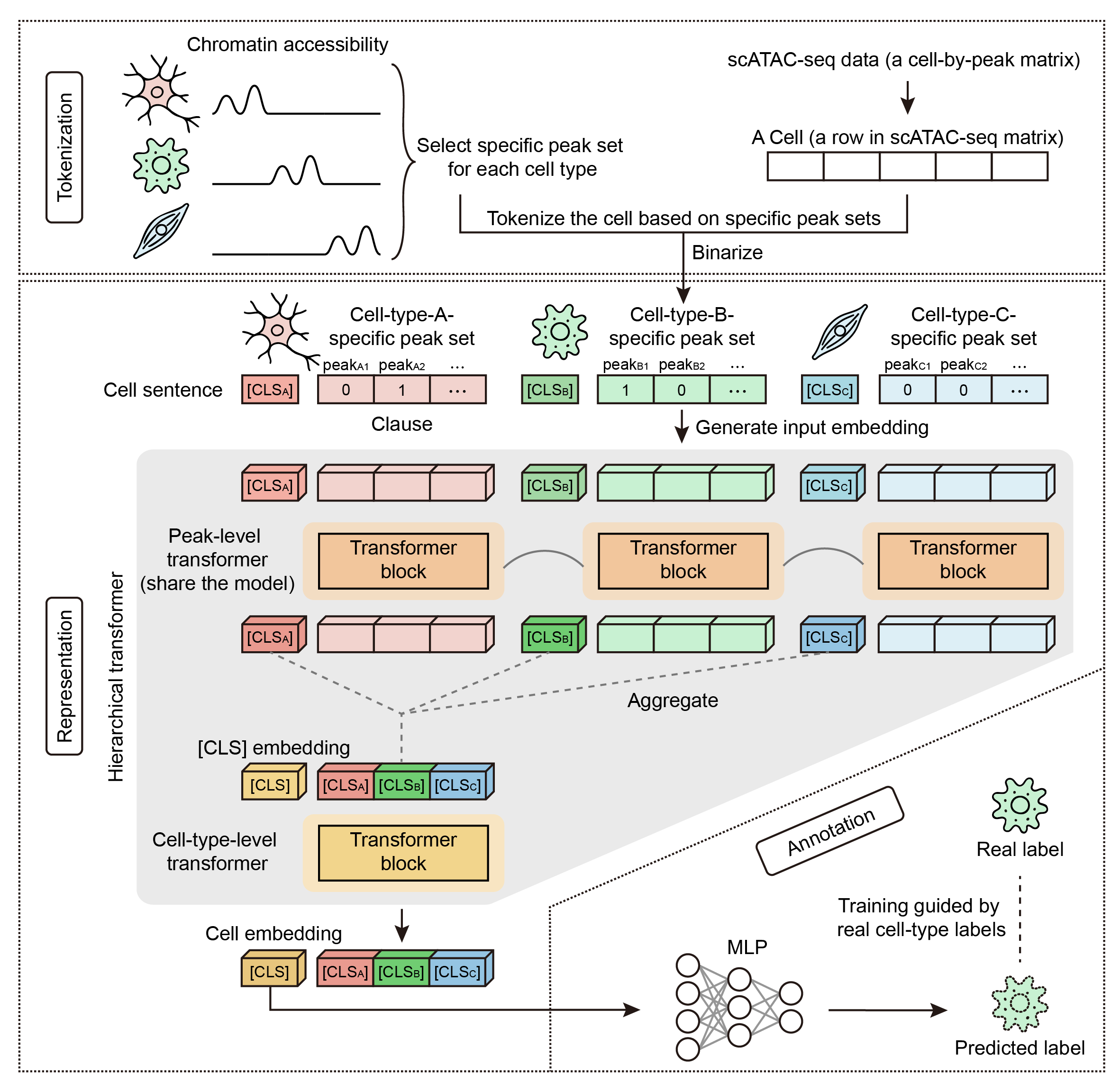
Extensive evaluations across scATAC-seq datasets demonstrate the robustness and high accuracy of HitAnno for intra-dataset annotation. In more practical inter-dataset annotation tasks, HitAnno effectively captures cellular heterogeneity despite batch effects, enabling superior performance. Leveraging this scalability, training on atlas-scale data empowers HitAnno to directly annotate new query datasets without retraining.
Online prediction
User Guide
Users can perform online cell type annotation using the HitAnno model on this website. First, users need to download the cCRE_hitanno.bed file (hg38). Then, follow the peaks in the file to transform and store scATAC-seq data in an h5ad file format (for detailed data processing code and file format, please refer to the Tutorial page). After selecting the h5ad file, users can click the Submit button to start the task. Optionally, users can provide an email address to receive notifications upon task completion. Once submitted, a task ID will be generated, which can be used to retrieve the status of the task later.